Groupwise Regularized Adaptive Sparse Precision Solution
The goal of grasps is to provide a collection of statistical methods that incorporate both element-wise and group-wise penalties to estimate a precision matrix, making them user-friendly and useful for researchers and practitioners.
\[ \hat{\Omega}(\lambda,\alpha,\gamma) = {\arg\min}_{\Omega \succ 0} \{ -\log\det(\Omega) + \text{tr}(S\Omega) + \mathcal{P}_{\lambda,\alpha,\gamma}(\Omega) \}, \] \[ \mathcal{P}_{\lambda,\alpha,\gamma}(\Omega) = \alpha \mathcal{P}^\text{idv}_{\lambda,\gamma}(\Omega) + (1-\alpha) \mathcal{P}^\text{grp}_{\lambda,\gamma}(\Omega), \] \[ \mathcal{P}^\text{idv}_{\lambda,\gamma}(\Omega) = \sum_{i,j} P_{\lambda,\gamma}(\lvert\omega_{ij}\rvert), \] \[ \mathcal{P}^\text{grp}_{\lambda,\gamma}(\Omega) = \sum_{g,g^\prime} P_{\lambda,\gamma}(\lVert\Omega_{gg^\prime}\rVert_F). \]
For more details, see the vignette Penalized Precision Matrix Estimation in grasps.
Penalties
The package grasps provides functions to estimate precision matrices using the following penalties:
| Penalty | Reference |
|---|---|
Lasso (penalty = "lasso") |
Tibshirani (1996); Friedman et al. (2008) |
Adaptive lasso (penalty = "adapt") |
Zou (2006); Fan et al. (2009) |
Atan (penalty = "atan") |
Wang and Zhu (2016) |
Exp (penalty = "exp") |
Wang et al. (2018) |
Lq (penalty = "lq") |
Frank and Friedman (1993); Fu (1998); Fan and Li (2001) |
LSP (penalty = "lsp") |
Candès et al. (2008) |
MCP (penalty = "mcp") |
Zhang (2010) |
SCAD (penalty = "scad") |
Fan and Li (2001); Fan et al. (2009) |
See the vignette Penalized Precision Matrix Estimation in grasps for more details.
Installation
- You can install the released version of grasps from CRAN with:
- You can install the development version of grasps from GitHub with:
Example
library(grasps)
## reproducibility for everything
set.seed(1234)
## block-structured precision matrix based on SBM
sim <- gen_prec_sbm(p = 30, K = 3,
within.prob = 0.25, between.prob = 0.05,
weight.dists = list("gamma", "unif"),
weight.paras = list(c(shape = 20, rate = 10),
c(min = 0, max = 5)),
cond.target = 100)
## ground truth visualization
plot(sim)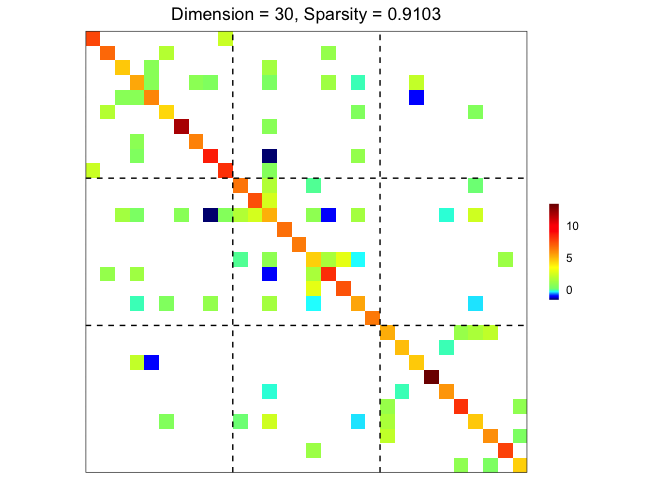
## n-by-p data matrix
library(MASS)
X <- mvrnorm(n = 20, mu = rep(0, 30), Sigma = sim$Sigma)
## precision matrix: adaptive lasso; BIC
prec <- grasps(X = X, membership = sim$membership, penalty = "adapt", crit = "BIC")
## precision matrix visualization
plot(prec)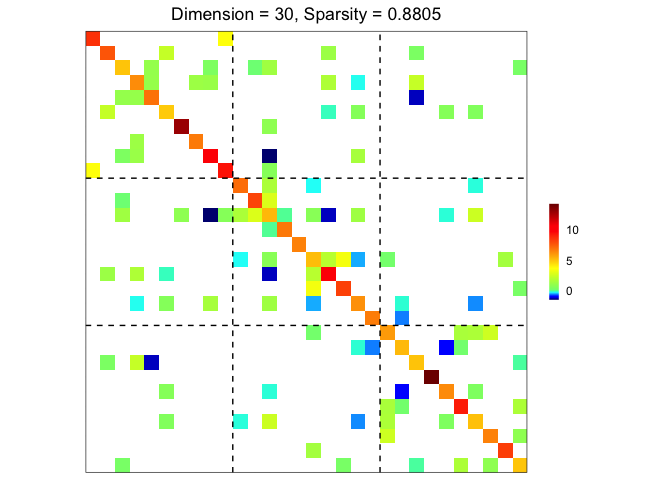
## performance
performance(hatOmega = prec$hatOmega, Omega = sim$Omega)
#> measure value
#> 1 sparsity 0.8805
#> 2 Frobenius 23.9013
#> 3 KL 7.6775
#> 4 quadratic 69.1639
#> 5 spectral 12.3571
#> 6 TP 23.0000
#> 7 TN 358.0000
#> 8 FP 29.0000
#> 9 FN 25.0000
#> 10 TPR 0.4792
#> 11 FPR 0.0749
#> 12 F1 0.4600
#> 13 MCC 0.3904
## adjacency matrix: diagonal = 0; raw partial correlations;
## no thresholding; weighted network
adj <- prec_to_adj(prec$hatOmega,
diag.zero = TRUE, absolute = FALSE,
threshold = NULL, weighted = TRUE)
## adjacency matrix visualization
plot(adj)